| Cat. # | Size | Qty. | Price |
|---|---|---|---|
| 9145T | 20 µl |
|
|
| 9145S | 100 µl |
|
|
| 9145L | 300 µl |
|
| REACTIVITY | H M R Mk |
| SENSITIVITY | Endogenous |
| MW (kDa) | 79, 86 |
| Source/Isotype | Rabbit IgG |
Product Information
For optimal ChIP and ChIP-seq results, use 5 μl of antibody and 10 μg of chromatin (approximately 4 x 106 cells) per IP. This antibody has been validated using SimpleChIP® Enzymatic Chromatin IP Kits.
| Application | Dilution |
|---|---|
| Western Blotting | 1:2000 |
| Simple Western™ | 1:10 - 1:50 |
| Immunoprecipitation | 1:100 |
| IHC Leica Bond | 1:100 - 1:400 |
| Immunohistochemistry (Paraffin) | 1:100 - 1:400 |
| Immunofluorescence (Immunocytochemistry) | 1:100 - 1:200 |
| Flow Cytometry (Fixed/Permeabilized) | 1:100 - 1:400 |
| Chromatin IP | 1:100 |
| Chromatin IP-seq | 1:100 |
For western blots, incubate membrane with diluted primary antibody in 5% w/v BSA, 1X TBS, 0.1% Tween® 20 at 4°C with gentle shaking, overnight.
NOTE: Please refer to primary antibody product webpage for recommended antibody dilution.
From sample preparation to detection, the reagents you need for your Western Blot are now in one convenient kit: #12957 Western Blotting Application Solutions Kit
NOTE: Prepare solutions with reverse osmosis deionized (RODI) or equivalent grade water.
Load 20 µl onto SDS-PAGE gel (10 cm x 10 cm).
NOTE: Loading of prestained molecular weight markers (#59329, 10 µl/lane) to verify electrotransfer and biotinylated protein ladder (#7727, 10 µl/lane) to determine molecular weights are recommended.
NOTE: Volumes are for 10 cm x 10 cm (100 cm2) of membrane; for different sized membranes, adjust volumes accordingly.
* Avoid repeated exposure to skin.
posted June 2005
revised June 2020
Protocol Id: 10
This protocol is intended for immunoprecipitation of native proteins utilizing Protein A agarose beads for analysis by western immunoblot or kinase activity.
NOTE: Prepare solutions with reverse osmosis deionized (RODI) or equivalent grade water.
10X Cell Lysis Buffer: (#9803) To prepare 10 ml of 1X cell lysis buffer, add 1 ml cell lysis buffer to 9 ml dH2O, mix.
NOTE: Add 1 mM PMSF (#8553) immediately prior to use.
IMPORTANT: Appropriate isotype controls are highly recommended in order to show specific binding in your primary antibody immunoprecipitation. Use Normal Rabbit IgG #2729 for rabbit polyclonal primary antibodies, Rabbit (DA1E) mAb IgG XP® Isotype Control #3900 for rabbit monoclonal primary antibodies, Mouse (G3A1) mAb IgG1 Isotype Control #5415 for mouse monoclonal IgG1 primary antibodies, Mouse (E5Y6Q) mAb IgG2a Isotype Control #61656 for mouse monoclonal IgG2a primary antibodies, Mouse (E7Q5L) mAb IgG2b Isotype Control #53484 for mouse monoclonal IgG2b primary antibodies, and Mouse (E1D5H) mAb IgG3 Isotype Control #37988 for mouse monoclonal IgG3 primary antibodies. Isotype controls should be concentration matched and run alongside the primary antibody samples.
Proceed to one of the following specific set of steps.
NOTE: When using primary antibodies produced in rabbit to detect proteins with a molecular weight in the range of 50 kDa, we recommend using Mouse Anti-Rabbit IgG (Light-Chain Specific) (D4W3E) mAb (#45262) or Mouse Anti-Rabbit IgG (Conformation Specific) (L27A9) mAb (#3678) (or HRP conjugate #5127) as a secondary antibody to minimize interference produced by denatured rabbit heavy chain. For proteins with a molecular weight in the range of 25 kDa, Mouse Anti-Rabbit IgG (Conformation Specific) (L27A9) mAb (#3678) (or HRP conjugate #5127) is recommended to minimize interference produced by denatured mouse light chain.
When using primary antibodies produced in mouse to detect proteins with a molecular weight in the range of 50 kDa, we recommend using Rabbit Anti-Mouse IgG (Light Chain Specific) (D3V2A) mAb (HRP Conjugate) (#58802) as a secondary antibody to minimize interference produced by denatured mouse heavy chain.
posted December 2008
revised October 2021
Protocol Id: 409
NOTE: Please see product datasheet or product webpage for appropriate antibody dilution^.
| Step | Reagents | Time/Temperature | |
|---|---|---|---|
| 1 | Dewax | BOND™ Dewax Solution, 100% Alcohol, BOND™ Wash Solution | Pre-programmed Leica® BOND™ |
| 2 | Antigen Retrieval | BOND™ Epitope Retrieval ER2 Solution | 20 min., 100˚C | Protocol: HIER 20 min with ER2 |
| 3 | Peroxide Block | Refine Detection Kit Peroxide Block* | 5 min. |
| WASH | BOND™ Wash Solution | 3x 0:00 min. | |
| 4 | Protein Block (optional) | #5425 NGS or #15019 Animal-Free Blocking Solution | 20 min. |
| 5 | Primary Antibody^ | Dilute in #8112 SignalStain® Antibody Diluent | 30 min. |
| WASH | BOND™ Wash Solution | 3x 2:00 min. | |
| NA | Post Primary Mouse Linker | Refine Detection Kit Post Primary* | Not Applied |
| 6 | Secondary Detection | Refine Detection Kit Polymer* | 10 min. |
| WASH | BOND™ Wash Solution/Deionized Water | Custom (see below) | |
| 7a | Visualization | Refine Detection Kit Mixed DAB Refine* | 0:00 min. |
| 7b | Visualization | Refine Detection Kit Mixed DAB Refine* | 10 min. |
| WASH | Deionized Water | 3x 0:00 min. | |
| 8 | Counterstain | Refine Detection Kit Hematoxylin* | 5 min. |
| WASH | Deionized Water | 0:00 min. | |
| WASH | BOND™ Wash Solution | 0:00 min. | |
| WASH | Deionized Water | 0:00 min. | |
| 9 | Dehydration (Offline): | ||
| Incubate sections in 95% ethanol two times for 10 seconds each. | |||
| Repeat in 100% ethanol, incubating sections two times for 10 seconds each. | |||
| Repeat in xylene, incubating sections two times for 10 seconds each. | |||
| 10 | Mount sections with coverslips and #14177 SignalStain® Mounting Medium | ||
| Optional Custom wash: | BOND™ Wash Solution | 2:00 | |
| BOND™ Wash Solution | Dispenser Type: OPEN 0:00 | ||
| BOND™ Wash Solution | 2:00 | ||
| BOND™ Wash Solution | Dispenser Type: OPEN 0:00 | ||
| BOND™ Wash Solution | 0:00 | ||
| Deionized Water | 0:00 | ||
*Reagent included in BOND™ Polymer Refine Detection Kit (Catalog No: DS9800)
LEICA® is a registered trademark of Leica Microsystems IR GmbH.
BOND™ is a trademark of Leica Biosystems Melbourne Pty. Ltd. No affiliation or sponsorship between CST and Leica Microsystems IR GmbH or Leica Biosystems Melbourne Pty. Ltd is implied.
posted August 2018
revised September 2018
Protocol Id: 1444
NOTE: Prepare solutions with reverse osmosis deionized (RODI) or equivalent grade water.
NOTE: Do not allow slides to dry at any time during this procedure.
For EDTA: Heat slides in a microwave submersed in 1X EDTA unmasking solution until boiling is initiated; follow with 15 min at a sub-boiling temperature (95°-98°C). No cooling is necessary.
| RECOMMENDED DETECTION REAGENTS |
SignalStain® Boost IHC Detection Reagent (HRP, Rabbit) #8114 | SignalStain® Boost IHC Detection Reagent (AP, Rabbit) #18653 |
|---|---|---|
|
COMPATIBLE CHROMOGEN |
SignalStain® DAB Substrate Kit #8059 | SignalStain® Vibrant Red Alkaline Phosphatase Substrate Kit #76713 |
| SignalStain® Vivid Purple Peroxidase Substrate Kit #96632 | SignalStain® Ultra Blue Alkaline Phosphatase Substrate Kit #12824 | |
| SignalStain® Deep Black Peroxidase Substrate Kit #72986 | ||
| SignalStain® Radiant Yellow Peroxidase Substrate Kit #69644 |
NOTE: Use of detection reagents other than those specified in this protocol may require further optimization of the primary antibody to account for the different sensitivities of the detection reagents.
posted February 2010
revised June 2020
Protocol Id: 284
NOTE: Prepare solutions with reverse osmosis deionized (RODI) or equivalently purified water.
Recommended Fluorochrome-conjugated Anti-Rabbit secondary antibodies:
NOTE: Cells should be grown, treated, fixed and stained directly in multiwell plates, chamber slides or on coverslips.
NOTE: All subsequent incubations should be carried out at room temperature unless otherwise noted in a humid light-tight box or covered dish/plate to prevent drying and fluorochrome fading.
posted November 2006
revised December 2010
Protocol Id: 32
All reagents required for this protocol may be efficiently purchased together in our Intracellular Flow Cytometry Kit (Methanol) #13593, or individually using the catalog numbers listed below.
NOTE: Prepare solutions with reverse osmosis deionized (RODI) or equivalent grade water.
NOTE: When including fluorescent cellular dyes in your experiment (including viability dyes, DNA dyes, etc.), please refer to the dye product page for the recommended protocol. Visit www.cellsignal.com for a full listing of cellular dyes validated for use in flow cytometry.
NOTE: Adherent cells or tissue should be dissociated and in single-cell suspension prior to fixation.
NOTE: Optimal centrifugation conditions will vary depending upon cell type and reagent volume. Generally, 150-300g for 1-5 minutes will be sufficient to pellet the cells.
NOTE: If using whole blood, lyse red blood cells and wash by centrifugation prior to fixation.
NOTE: Antibodies targeting CD markers or other extracellular proteins may be added prior to fixation if the epitope is disrupted by formaldehyde and/or methanol. The antibodies will remain bound to the target of interest during the fixation and permeabilization process. However, note that some fluorophores (including PE and APC) are damaged by methanol and thus should not be added prior to permeabilization. Conduct a small-scale experiment if you are unsure.
NOTE: Count cells using a hemocytometer or alternative method.
posted July 2009
revised June 2020
Protocol Id: 404
Specific for product: SimpleChIP® Plus Sonication Chromatin IP Kit #56383.
Reagents Included:
Reagents Not Included:
| ! | This ! signifies an important step in the protocol regarding volume changes based on the number of immunoprecipitation preparations (IP preps). One IP prep is defined as 4 x 106 tissue cultured cells or 25 mg or disaggregated tissue. |
| !! | This !! signifies an important step to dilute a buffer before proceeding. |
| SAFE STOP | This is a safe stopping point in the protocol, if stopping is necessary. |
When harvesting tissue, remove unwanted material from the sample, such as fat and necrotic material. Tissue can be processed and cross-linked immediately or frozen on dry ice for processing later. For optimal ChIP results, use 25 mg of tissue for each immunoprecipitation to be performed. An additional 5 mg of tissue should be processed for Analysis of Chromatin Digestion and Concentration and for use as input chromatin (Section IV). The chromatin yield does vary between tissue types, and some tissues may require more than 25 mg for each immunoprecipitation.
One chromatin preparation is defined as 100 to 150 mg of tissue. This recommended amount of tissue accounts for potential low yield with some tissue types and also ensures efficient chromatin fragmentation during sonication. Please see Appendix A for more information regarding the expected chromatin yield for different types of tissue.
Before starting:
(!) All buffer volumes should be increased proportionally based on the number of chromatin preps in the experiment.
For optimal ChIP results, use approximately 4 x 106 cells for each immunoprecipitation to be performed. For HCT 116 cells, this is equivalent to 1/3 of a 15 cm culture dish containing cells that are 90% confluent in 20 ml of growth medium. An additional 1 x 106 cells should be processed for Analysis of Chromatin Digestion and Concentration and for use as input chromatin (Section IV).
One chromatin preparation is defined as 1 x 107 to 2 x 107 cells. This recommended cell number accounts for potential low yield with some cell types and also ensures efficient chromatin fragmentation during sonication.
Before starting:
(!) All buffer volumes should be increased proportionally based on the number of 15 cm tissue culture dishes (or 20 ml suspension cells) used in the experiment.
One chromatin preparation is defined as 100 to 150 mg of tissue or 1 x 107to 2 x 107tissue culture cells. Multiple chromatin preparations can be performed simultaneously, as long as the amounts of buffers are scaled appropriately and sonication is performed on 1 ml samples. The number of cells and volume of sample used for sonication is critical for generation of appropriately sized chromatin fragments.
Before starting:
(!) All buffer volumes should be increased proportionally based on the number of chromatin preps in the experiment.
For optimal ChIP results, use approximately 5 to 10 µg of sonicated, cross-linked chromatin (as determined in Section IV) per immunoprecipitation. This should be roughly equivalent to a single 100 µl IP prep from 25 mg of disaggregated tissue or 4x106 tissue culture cells. Typically, 100 µl of digested chromatin is diluted into 400 µl 1X ChIP Buffer prior to the addition of antibodies. However, if more than 100 µl of chromatin is required per IP, the cross-linked chromatin preparation must be diluted into 1X ChIP buffer at a dilution ratio of 1:4. No additional protein G magnetic beads are necessary in this case, although prolonged incubation time with beads is helpful.
Before starting:
(!) All buffer volumes should be increased proportionally based on the number of immunoprecipitations in the experiment.
NOTE: Most antibodies from Cell Signaling Technology work optimally between 1 and 2 µg per IP sample. In the case where there are multiple samples with varying concentrations, it is best to match the negative control Normal Rabbit IgG #2729 to the highest antibody concentration.
Before starting:
(!) All buffer volumes should be increased proportionally based on the number of immunoprecipitations in the experiment.
Before starting:
Recommendations:
| Primer length: | 24 nucleotides |
| Optimum Tm: | 60°C |
| Optimum GC: | 50% |
| Amplicon size: | 150 to 200 bp (for standard PCR) |
| 80 to 160 bp (for real-time quantitative PCR) |
Standard PCR Method:
| Reagent | Volume for 1 PCR Reaction (18 µl) |
|---|---|
| Nuclease-free H2O | 12.5 µl |
| 10X PCR Buffer | 2.0 µl |
| 4 mM dNTP Mix | 1.0 µl |
| 5 µM RPL30 Primers | 2.0 µl |
| Taq DNA Polymerase | 0.5 µl |
Real-Time Quantitative PCR Method:
| Reagent | Volume for 1 PCR Reaction (18 µl) |
|---|---|
| Nuclease-free H2O | 6 µl |
| 5 µM RPL30 Primers | 2 µl |
| SimpleChIP® Universal qPCR Master Mix #88989 | 10 µl |
| a. | Initial Denaturation | 95°C 3 min |
| b. | Denature | 95°C 15 sec |
| c. | Anneal and Extension: | 60°C 60 sec |
| d. | Repeat steps b and c for a total of 40 cycles. |
Percent Input = 2% x 2(C[T] 2%Input Sample - C[T] IP Sample)
C[T] = CT = Average threshold cycle of PCR reaction
The immuno-enriched DNA samples prepared with this kit are directly compatible with ChIP-seq. For downstream NG-sequencing DNA library construction, use a DNA library preparation protocol or kit compatible with your downstream sequencing platform. For sequencing on Illumina® platforms, we recommend DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795 and its associated index primers Multiplex Oligos for Illumina® (Single Index Primers) (ChIP-seq, CUT&RUN) #29580 or Multiplex Oligos for Illumina® (Dual Index Primers) (ChIP-seq, CUT&RUN) #47538.
Recommendations:
When harvesting cross-linked chromatin from tissue samples, the yield of chromatin can vary significantly between tissue types. The table to the right provides a range for the expected yield of chromatin from 100 mg of tissue compared to 2 x 107 HCT 116 cells, and the expected DNA concentration, as determined in Section IV of the protocol. For optimal ChIP results, we recommend DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795 and its associated index primers Multiplex Oligos for Illumina® (Single Index Primers) (ChIP-seq, CUT&RUN) #29580 or Multiplex Oligos for Illumina® (Dual Index Primers) (ChIP-seq, CUT&RUN) #47538.
| Tissue/Cell | Total Chromatin Yield | Expected DNA Concentration |
|---|---|---|
| Liver | 50 µg per 100 mg tissue | 150 µg/ml |
| Brain | 25 µg per 100 mg tissue | 50 µg/ml |
| Heart | 105 µg per 100 mg tissue | 20 µg/ml |
| HCT 116 | 100-150 µg per 2 x 107 cells | 100-150 µg/ml |
Transcription factors and cofactors bind to chromatin DNA more loosely than histone proteins. As a result, they tend to dissociate from chromatin during sonication. Additional fixation time can result in increased capture of transcription factors and cofactors in the ChIP assay, especially with tissue samples. As shown in Figure 7, increasing the fixation time from 10 min to 30 min may reduce chromatin fragmentation size (left panel), but it significantly enhances enrichment of both cofactors RING1B and SUZ12 in heart tissue, as indicated by ChIP-qPCR (middle and right panels).
Typically, 10 min fixation is sufficient for histone modification ChIP with both cell and tissue samples, whereas transcription factors and cofactors may require additional fixation up to 30 min, especially with tissue samples.
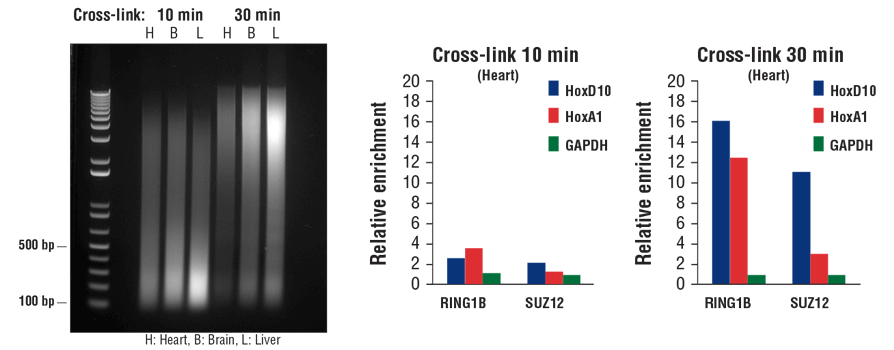
FIGURE 7. Mouse heart (H), brain (B), and liver (L) were cross-linked for 10 min or 30 min, as indicated (left panel). The chromatin was prepared and sonicated, DNA was purified and separated by electrophoresis on a 1% agarose gel. In the ChIP-qPCR assay (middle and right panels), chromatin immunoprecipitations were performed with either RING1B (D22F2) XP® Rabbit mAb #5694 or SUZ12 (D39F6) XP® Rabbit mAb #3737. The enriched DNA was quantified by real-time PCR using SimpleChIP® Mouse HoxD10 Exon 1 Primers #7429, SimpleChIP® Mouse HoxA1 Promoter Primers #7341, and SimpleChIP® Mouse GAPDH Intron 2 Primers #8986. The amount of immunoprecipitated DNA in each sample is represented as normalized signal to the negative GAPDH locus (equivalent to one).
Optimal conditions for the fragmentation of cross-linked chromatin DNA is highly dependent on the number of cells, volume of sample, length of sonication, and sonicator power setting used. For each sonication sample, we recommend using 100-150 mg of tissue or 1 x 107 to 2 x 107 cells per 1 ml ChIP Sonication Nuclear Lysis Buffer. Below is a protocol for determination of the optimal sonication conditions for a specific tissue or cell type.
NOTE: Optimal sonication conditions can vary with different sample types and fixation times. Use the minimal number of sonication cycles required to generate the desired length of chromatin fragments. Over sonication, as indicated by more than 80% of total DNA fragments less than 500 bp, can result in excessive damage to the chromatin and result in lower immunoprecipitation efficiency (see Figure 8, right panel).
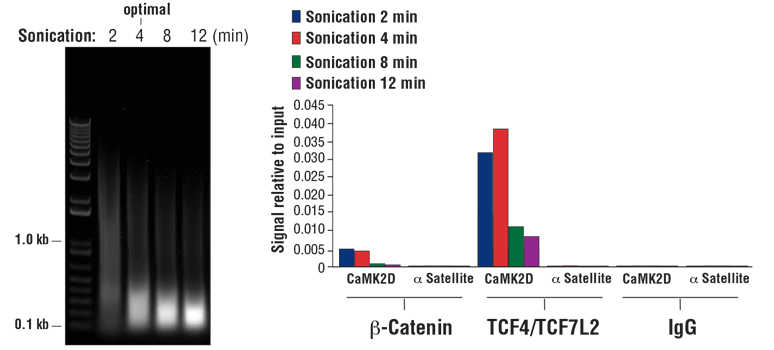
FIGURE 8. HCT 116 cells were cross-linked for 10 min and sonicated for the time indicated (left panel). DNA was purified as described in Section IV of the SimpleChIP® Plus Sonication Chromatin IP Kit #56383, and 20 µl of purified DNA was separated by electrophoresis on a 1% agarose gel. As shown in the left panel, increasing cycles of sonication reduces the size of chromatin fragments. Chromatin immunoprecipitations were performed with either Non-phospho (Active) Β-Catenin (Ser33/37/Thr41) (D13A1) Rabbit mAb #8814, TCF4/TCF7L2 (C48H11) Rabbit mAb #2569, or Normal Rabbit IgG #2729 using SimpleChIP® Plus Sonication Chromatin IP Kit #56383. The enriched DNA was quantified by real-time PCR using SimpleChIP® Human CaMK2D Intron 3 Primers #5111 and SimpleChIP® Human α Satellite Repeat Primers #4486. The amount of immunoprecipitated DNA in each sample is represented as signal relative to the total amount of input chromatin (equivalent to 100%; right panel). As shown, 4 min of chromatin sonication is optimal when using a Branson Digital Sonifier D250 probe sonicator with a 1/8 inch Micro Tip. Over-sonication significantly impairs enrichment of both cofactor beta-catenin and transcription factor TCF4/TCF7L2 containing chromatin.
| Problem | Possible Causes | Recommendation |
|---|---|---|
| 1. Concentration of the fragmented chromatin is too low. |
Cell/nuclear lysis is incomplete. Not enough cells were used for the chromatin preparation. |
If DNA concentration of the chromatin preparation is close to 50 µg/ml, add additional chromatin to each IP to give at least 5 µg/IP and continue with protocol. Count a separate plate of cells before cross-linking to determine an accurate cell number. |
| 2. Chromatin is under-fragmented and fragments are too large (>50% above 1.5kb). |
Cells may have been over cross-linked. Too many cells/tissues were processed. |
Shorten the crosslinking time within 10-30 minute range. Reduce the amount of cell/tissues per sonication. Conduct sonication time course. |
| 3. Chromatin is over-fragmented (>90% under 300 bp). |
Sonication condition is too harsh. |
Conduct a sonication time course to find a minimum output/duration to achieve appropriate sonication. |
| 4. No product or very little product in the input PCR reactions. |
Not enough DNA added to the PCR reaction or conditions are not optimal. PCR amplified region may span nucleosome-free region. Not enough chromatin added to the IP or chromatin is over-sonicated |
Add more DNA to the PCR reaction or increase the number of amplification cycles. Optimize the PCR conditions for experimental primer set using purified DNA from cross-linked and sonicated chromatin. For optimal ChIP results add 5-10 µg chromatin per IP. See recommendations for problems 1 and 3 above. |
| 5. No product in the positive control Histone H3-IP RPL30 PCR reaction. |
Not enough chromatin or antibody added to the IP reaction or IP incubation time is too short. Incomplete elution of chromatin from Protein G beads. |
Be sure to add 5-10 µg of chromatin and 10 µl of antibody to each IP reaction and incubate with antibody over-night and an additional 2 hr after adding Protein G beads. Elution of chromatin from Protein G beads is optimal at 65°C with frequent mixing to keep beads suspended in solution. |
| 6. Quantity of product in the negative control Rabbit IgG-IP and positive control Histone H3-IP PCR reactions is equivalent. |
Too much or not enough chromatin added to the IP reaction. Alternatively, too much antibody added to the IP reaction. Too much DNA added to the PCR reaction or too many cycles of amplification. |
Add no more than 15 µg of chromatin and 10 µl of histone H3 antibody to each IP reaction. Reduce the amount of normal rabbit IgG to 1 µl per IP. Add less DNA to the PCR reaction or decrease the number of PCR cycles. It is very important that the PCR products are analyzed within the linear amplification phase of PCR. Otherwise, the differences in quantities of starting DNA cannot be accurately measured. |
| 7. No product in the Experimental Antibody-IP PCR reaction. |
Not enough DNA added to the PCR reaction. Not enough antibody added to the IP reaction. Antibody does not work for IP. |
Add more DNA to the PCR reaction or increase the number of amplification cycles. Typically a range of 1 to 5 µg of antibody is added to the IP reaction; however, the exact amount depends greatly on the individual antibody. Increase the amount of antibody added to the IP. Find an alternate antibody source. |
posted March 2017
revised April 2022
Protocol Id: 1384
Specific for product: SimpleChIP® Plus Sonication Chromatin IP Kit #56383.
When harvesting tissue, remove unwanted material from the sample, such as fat and necrotic material. Tissue can be processed and cross-linked immediately or frozen on dry ice for processing later. For optimal ChIP results, use 25 mg of tissue for each immunoprecipitation to be performed. An additional 5 mg of tissue should be processed for Analysis of Chromatin Digestion and Concentration and for use as input chromatin (Section IV). The chromatin yield does vary between tissue types, and some tissues may require more than 25 mg for each immunoprecipitation.
One chromatin preparation is defined as 100 to 150 mg of tissue. This recommended amount of tissue accounts for potential low yield with some tissue types and also ensures efficient chromatin fragmentation during sonication. Please see Appendix A for more information regarding the expected chromatin yield for different types of tissue.
Before starting:
For optimal ChIP results, use approximately 4 x 106 cells for each immunoprecipitation to be performed. For HCT 116 cells, this is equivalent to 1/3 of a 15 cm culture dish containing cells that are 90% confluent in 20 ml of growth medium. An additional 1 x 106 cells should be processed for Analysis of Chromatin Digestion and Concentration and for use as input chromatin (Section IV).
One chromatin preparation is defined as 1 x 107 to 2 x 107 cells. This recommended cell number accounts for potential low yield with some cell types and also ensures efficient chromatin fragmentation during sonication.
Before starting:
One chromatin preparation is defined as 100 to 150 mg of tissue or 1 x 107to 2 x 107tissue culture cells. Multiple chromatin preparations can be performed simultaneously, as long as the amounts of buffers are scaled appropriately and sonication is performed on 1 ml samples. The number of cells and volume of sample used for sonication is critical for generation of appropriately sized chromatin fragments.
Before starting:
For optimal ChIP results, use approximately 5 to 10 µg of sonicated, cross-linked chromatin (as determined in Section IV) per immunoprecipitation. This should be roughly equivalent to a single 100 µl IP prep from 25 mg of disaggregated tissue or 4x106 tissue culture cells. Typically, 100 µl of digested chromatin is diluted into 400 µl 1X ChIP Buffer prior to the addition of antibodies. However, if more than 100 µl of chromatin is required per IP, the cross-linked chromatin preparation must be diluted into 1X ChIP buffer at a dilution ratio of 1:4. No additional protein G magnetic beads are necessary in this case, although prolonged incubation time with beads is helpful.
Before starting:
NOTE: Most antibodies from Cell Signaling Technology work optimally between 1 and 2 µg per IP sample. In the case where there are multiple samples with varying concentrations, it is best to match the negative control Normal Rabbit IgG #2729 to the highest antibody concentration.
Before starting:
Before starting:
Recommendations:
| Primer length: | 24 nucleotides |
| Optimum Tm: | 60°C |
| Optimum GC: | 50% |
| Amplicon size: | 150 to 200 bp (for standard PCR) |
| 80 to 160 bp (for real-time quantitative PCR) |
Standard PCR Method:
| Reagent | Volume for 1 PCR Reaction (18 µl) |
|---|---|
| Nuclease-free H2O | 12.5 µl |
| 10X PCR Buffer | 2.0 µl |
| 4 mM dNTP Mix | 1.0 µl |
| 5 µM RPL30 Primers | 2.0 µl |
| Taq DNA Polymerase | 0.5 µl |
Real-Time Quantitative PCR Method:
| Nuclease-free H2O | 6 µl |
| 5 µM RPL30 Primers | 2 µl |
| SimpleChIP® Universal qPCR Master Mix #88989 | 10 µl |
| a. | Initial Denaturation | 95°C 3 min |
| b. | Denature | 95°C 15 sec |
| c. | Anneal and Extension: | 60°C 60 sec |
| d. | Repeat steps b and c for a total of 40 cycles. |
Percent Input = 2% x 2(C[T] 2%Input Sample - C[T] IP Sample)
C[T] = CT = Average threshold cycle of PCR reaction
The immuno-enriched DNA samples prepared with this kit are directly compatible with ChIP-seq. For downstream NG-sequencing DNA library construction, use a DNA library preparation protocol or kit compatible with your downstream sequencing platform. For sequencing on Illumina® platforms, we recommend DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795 and its associated index primers Multiplex Oligos for Illumina® (Single Index Primers) (ChIP-seq, CUT&RUN) #29580 or Multiplex Oligos for Illumina® (Dual Index Primers) (ChIP-seq, CUT&RUN) #47538.
Recommendations:
When harvesting cross-linked chromatin from tissue samples, the yield of chromatin can vary significantly between tissue types. The table to the right provides a range for the expected yield of chromatin from 100 mg of tissue compared to 2 x 107 HCT 116 cells, and the expected DNA concentration, as determined in Section IV of the protocol. For optimal ChIP results, we recommend DNA Library Prep Kit for Illumina® (ChIP-seq, CUT&RUN) #56795 and its associated index primers Multiplex Oligos for Illumina® (Single Index Primers) (ChIP-seq, CUT&RUN) #29580 or Multiplex Oligos for Illumina® (Dual Index Primers) (ChIP-seq, CUT&RUN) #47538.
| Tissue/Cell | Total Chromatin Yield | Expected DNA Concentration |
|---|---|---|
| Liver | 50 µg per 100 mg tissue | 150 µg/ml |
| Brain | 25 µg per 100 mg tissue | 50 µg/ml |
| Heart | 105 µg per 100 mg tissue | 20 µg/ml |
| HCT 116 | 100-150 µg per 2 x 107 cells | 100-150 µg/ml |
Transcription factors and cofactors bind to chromatin DNA more loosely than histone proteins. As a result, they tend to dissociate from chromatin during sonication. Additional fixation time can result in increased capture of transcription factors and cofactors in the ChIP assay, especially with tissue samples. As shown in Figure 7, increasing the fixation time from 10 min to 30 min may reduce chromatin fragmentation size (left panel), but it significantly enhances enrichment of both cofactors RING1B and SUZ12 in heart tissue, as indicated by ChIP-qPCR (middle and right panels).
Typically, 10 min fixation is sufficient for histone modification ChIP with both cell and tissue samples, whereas transcription factors and cofactors may require additional fixation up to 30 min, especially with tissue samples.

FIGURE 7. Mouse heart (H), brain (B), and liver (L) were cross-linked for 10 min or 30 min, as indicated (left panel). The chromatin was prepared and sonicated, DNA was purified and 20 µl was separated by electrophoresis on a 1% agarose gel. In the ChIP-qPCR assay (middle and right panels), chromatin immunoprecipitations were performed with either 10 µl of RING1B (D22F2) XP® Rabbit mAb #5694 or 5 µl of SUZ12 (D39F6) XP® Rabbit mAb #3737. The enriched DNA was quantified by real-time PCR using SimpleChIP® Mouse HoxD10 Exon 1 Primers #7429, SimpleChIP® Mouse HoxA1 Promoter Primers #7341, and SimpleChIP® Mouse GAPDH Intron 2 Primers #8986. The amount of immunoprecipitated DNA in each sample is represented as normalized signal to the negative GAPDH locus (equivalent to one).
Optimal conditions for the fragmentation of cross-linked chromatin DNA is highly dependent on the number of cells, volume of sample, length of sonication, and sonicator power setting used. For each sonication sample, we recommend using 100-150 mg of tissue or 1 x 107 to 2 x 107 cells per 1 ml ChIP Sonication Nuclear Lysis Buffer. Below is a protocol for determination of the optimal sonication conditions for a specific tissue or cell type.
NOTE: Optimal sonication conditions can vary with different sample types and fixation times. Use the minimal number of sonication cycles required to generate the desired length of chromatin fragments. Over sonication, as indicated by more than 80% of total DNA fragments less than 500 bp, can result in excessive damage to the chromatin and result in lower immunoprecipitation efficiency (see Figure 8, right panel).

FIGURE 8. Chromatin immunoprecipitations were performed with 2 x 107 HCT 116 cells that were cross-linked for 10 min and sonicated for the time indicated (left panel). DNA was purified as described in Section IV of the SimpleChIP® Plus Sonication Chromatin IP Kit #56383, and 20 µl of purified DNA was separated by electrophoresis on a 1% agarose gel. As shown in the left panel, increasing cycles of sonication reduces the size of chromatin fragments. Chromatin immunoprecipitations were performed with either 5 µl of Non-phospho (Active) Β-Catenin (Ser33/37/Thr41) (D13A1) Rabbit mAb #8814, 10 µl of TCF4/TCF7L2 (C48H11) Rabbit mAb #2569, or 2 µl of Normal Rabbit IgG #2729 using SimpleChIP® Plus Sonication Chromatin IP Kit #56383. The enriched DNA was quantified by real-time PCR using SimpleChIP® Human CaMK2D Intron 3 Primers #5111 and SimpleChIP® Human α Satellite Repeat Primers #4486. The amount of immunoprecipitated DNA in each sample is represented as signal relative to the total amount of input chromatin (equivalent to one; right panel). As shown, 4 min of chromatin sonication is optimal when using a Branson Digital Sonifier D250 probe sonicator with a 1/8 inch Micro Tip. Over-sonication significantly impairs enrichment of both cofactor beta-catenin and transcription factor TCF4/TCF7L2 containing chromatin.
| Problem | Possible Causes | Recommendation |
|---|---|---|
| 1. Concentration of the fragmented chromatin is too low. |
Cell/nuclear lysis is incomplete. Not enough cells were used for the chromatin preparation. |
If DNA concentration of the chromatin preparation is close to 50 µg/ml, add additional chromatin to each IP to give at least 5 µg/IP and continue with protocol. Count a separate plate of cells before cross-linking to determine an accurate cell number. |
| 2. Chromatin is under-fragmented and fragments are too large (>50% above 1.5kb). |
Cells may have been over cross-linked. Too many cells/tissues were processed. |
Shorten the crosslinking time within 10-30 minute range. Reduce the amount of cell/tissues per sonication. Conduct sonication time course. |
| 3. Chromatin is over-fragmented (>90% under 300 bp). |
Sonication condition is too harsh. |
Conduct a sonication time course to find a minimum output/duration to achieve appropriate sonication. |
| 4. No product or very little product in the input PCR reactions. |
Not enough DNA added to the PCR reaction or conditions are not optimal. PCR amplified region may span nucleosome-free region. Not enough chromatin added to the IP or chromatin is over-sonicated |
Add more DNA to the PCR reaction or increase the number of amplification cycles. Optimize the PCR conditions for experimental primer set using purified DNA from cross-linked and sonicated chromatin. For optimal ChIP results add 5-10 µg chromatin per IP. See recommendations for problems 1 and 3 above. |
| 5. No product in the positive control Histone H3-IP RPL30 PCR reaction. |
Not enough chromatin or antibody added to the IP reaction or IP incubation time is too short. Incomplete elution of chromatin from Protein G beads. |
Be sure to add 5-10 µg of chromatin and 10 µl of antibody to each IP reaction and incubate with antibody over-night and an additional 2 hr after adding Protein G beads. Elution of chromatin from Protein G beads is optimal at 65°C with frequent mixing to keep beads suspended in solution. |
| 6. Quantity of product in the negative control Rabbit IgG-IP and positive control Histone H3-IP PCR reactions is equivalent. |
Too much or not enough chromatin added to the IP reaction. Alternatively, too much antibody added to the IP reaction. Too much DNA added to the PCR reaction or too many cycles of amplification. |
Add no more than 15 µg of chromatin and 10 µl of histone H3 antibody to each IP reaction. Reduce the amount of normal rabbit IgG to 1 µl per IP. Add less DNA to the PCR reaction or decrease the number of PCR cycles. It is very important that the PCR products are analyzed within the linear amplification phase of PCR. Otherwise, the differences in quantities of starting DNA cannot be accurately measured. |
| 7. No product in the Experimental Antibody-IP PCR reaction. |
Not enough DNA added to the PCR reaction. Not enough antibody added to the IP reaction. Antibody does not work for IP. |
Add more DNA to the PCR reaction or increase the number of amplification cycles. Typically a range of 1 to 5 µg of antibody is added to the IP reaction; however, the exact amount depends greatly on the individual antibody. Increase the amount of antibody added to the IP. Find an alternate antibody source. |
posted March 2017
revised April 2022
Protocol Id: 1404
Human, Mouse, Rat, Monkey
Hamster, Bovine, Pig, Horse
Monoclonal antibody is produced by immunizing animals with a synthetic phosphopeptide corresponding to residues surrounding Tyr705 of mouse Stat3.
The Stat3 transcription factor is an important signaling molecule for many cytokines and growth factor receptors (1) and is required for murine fetal development (2). Research studies have shown that Stat3 is constitutively activated in a number of human tumors (3,4) and possesses oncogenic potential (5) and anti-apoptotic activities (3). Stat3 is activated by phosphorylation at Tyr705, which induces dimerization, nuclear translocation, and DNA binding (6,7). Transcriptional activation seems to be regulated by phosphorylation at Ser727 through the MAPK or mTOR pathways (8,9). Stat3 isoform expression appears to reflect biological function as the relative expression levels of Stat3α (86 kDa) and Stat3β (79 kDa) depend on cell type, ligand exposure, or cell maturation stage (10). It is notable that Stat3β lacks the serine phosphorylation site within the carboxy-terminal transcriptional activation domain (8).
Explore pathways related to this product.
Except as otherwise expressly agreed in a writing signed by a legally authorized representative of CST, the following terms apply to Products provided by CST, its affiliates or its distributors. Any Customer's terms and conditions that are in addition to, or different from, those contained herein, unless separately accepted in writing by a legally authorized representative of CST, are rejected and are of no force or effect.
Products are labeled with For Research Use Only or a similar labeling statement and have not been approved, cleared, or licensed by the FDA or other regulatory foreign or domestic entity, for any purpose. Customer shall not use any Product for any diagnostic or therapeutic purpose, or otherwise in any manner that conflicts with its labeling statement. Products sold or licensed by CST are provided for Customer as the end-user and solely for research and development uses. Any use of Product for diagnostic, prophylactic or therapeutic purposes, or any purchase of Product for resale (alone or as a component) or other commercial purpose, requires a separate license from CST. Customer shall (a) not sell, license, loan, donate or otherwise transfer or make available any Product to any third party, whether alone or in combination with other materials, or use the Products to manufacture any commercial products, (b) not copy, modify, reverse engineer, decompile, disassemble or otherwise attempt to discover the underlying structure or technology of the Products, or use the Products for the purpose of developing any products or services that would compete with CST products or services, (c) not alter or remove from the Products any trademarks, trade names, logos, patent or copyright notices or markings, (d) use the Products solely in accordance with CST Product Terms of Sale and any applicable documentation, and (e) comply with any license, terms of service or similar agreement with respect to any third party products or services used by Customer in connection with the Products.